Macro biomolecules
From SMOSwiki
Introduction
Our goal is to understand the molecular mechanism of working macro biomolecules such as DNA, lipid, and protein. A simple way to do this is 'whaiching' the biomolecules in action in real time. Biomolecules are, unfortunately, too small to directly observe. To overcome this limitation, we exploit the merit of fluorescence techniques, such as time-resolved fluorescence spectroscopy (see TCSPC), fluorescence resonance energy transfer (FRET) or fluorescence correlation spectroscopy (FCS), which are non-invasive proving tools. That is, we attach fluorescence tag on the molecules and let them do their job, so that we can simply watch them. The labeled fluorescenct dyes on the target molecules would give a variety information including high prescision temporal and spacial movement and intra molecular motion of the molecules. With this simple approach, we attempt to give answers on the open questions in biology such as the conformational change of proteins, the biological role of nucleic acid structure, and microscopic domain formation of lipids bilayer.
RecA filament dynamics
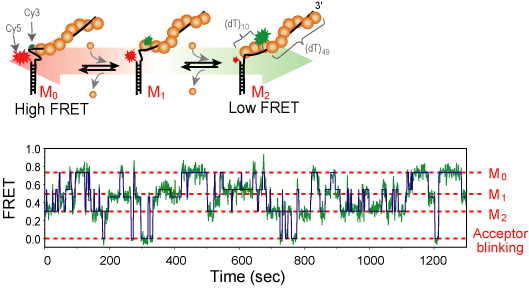
RecA is a protein involved in several DNA repair pathways s.a. homologous recombination and SOS response. RecA forms a right handed helical filament on a single-strand DNA, so that the DNA can be stretched out allowing the homologous sequence searching [1]. We use single-molecule FRET technique to observe the formation and growth of RecA filament in real-time. we could resolve the monomer binding and dissociation from the filament and get the kinetic rate under various chemical conditions.[2]
We labele single stranded DNA with Cy3 and Cy5 dye as a FRET donor and acceptor, respectively. The dyes are typically 5~13 nucleotide apart so that a few RecA monomers can bind between them. As the RecA stretches the DNA about 1.5 fold in length, the inter-dye distance change can be monitored by FRET in real-time. From the measured FRET time trajectory under the various chemical condition, we could calculate the biding and dissociation rate. From the result, we expect to elucidate the molecular mechanism of RecA filament formation and degradation.
DNA bubble and fork formation
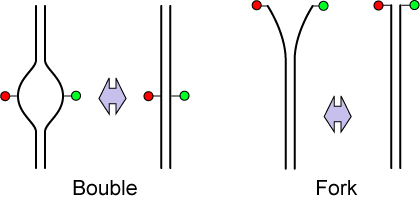
DNA has a double-helical structure with two complimentary strand. In the aqueous solution, however, partial unwinding of double-strand DNA happens spontaneously by thermal (or quantum) fluctuation, which is called DNA bubble and fork formation. As FRET is sensitive to the distance change in nanometer scale, we study the DNA bubble and fork formation in real time[3]. We also use the Fluorescence Correlation Spectroscopy technique, which is useful to extract out the hidden information in the noisy signal.[4]. We examine the bubble and fork dynamics by changing the salt concentration in the solution.
References:
- ↑ Bell, C. E. (2005). Structure and Mechanism of Escherichia coli RecA ATPase. Mol Microbiol 58, 358-366.
- ↑ Joo, C., McKinney, S. A., Nakamura, M., Rasnik, I., Myoung, S., and Ha, T. (2006). Real-Time Observation of RecA Filament Dynamics with Single Monomer Resolution. Cell, 126, 515-527.
- ↑ Yuan, C., Chen, H., Lou, X. W., and Archer, L. A. (2008). DNA Bending Stiffness on Small Length Scales. Phys. Rev. Lett., 100, 18102.
- ↑ Altan-Bonnet, G., Libchaber, A., and Krichevsky, O. (2003) Bubble dynamics in Double-Stranded DNA. Phys. Rev. Lett., 90, 138101.